What’s new in R 4.6.0? | R-bloggers
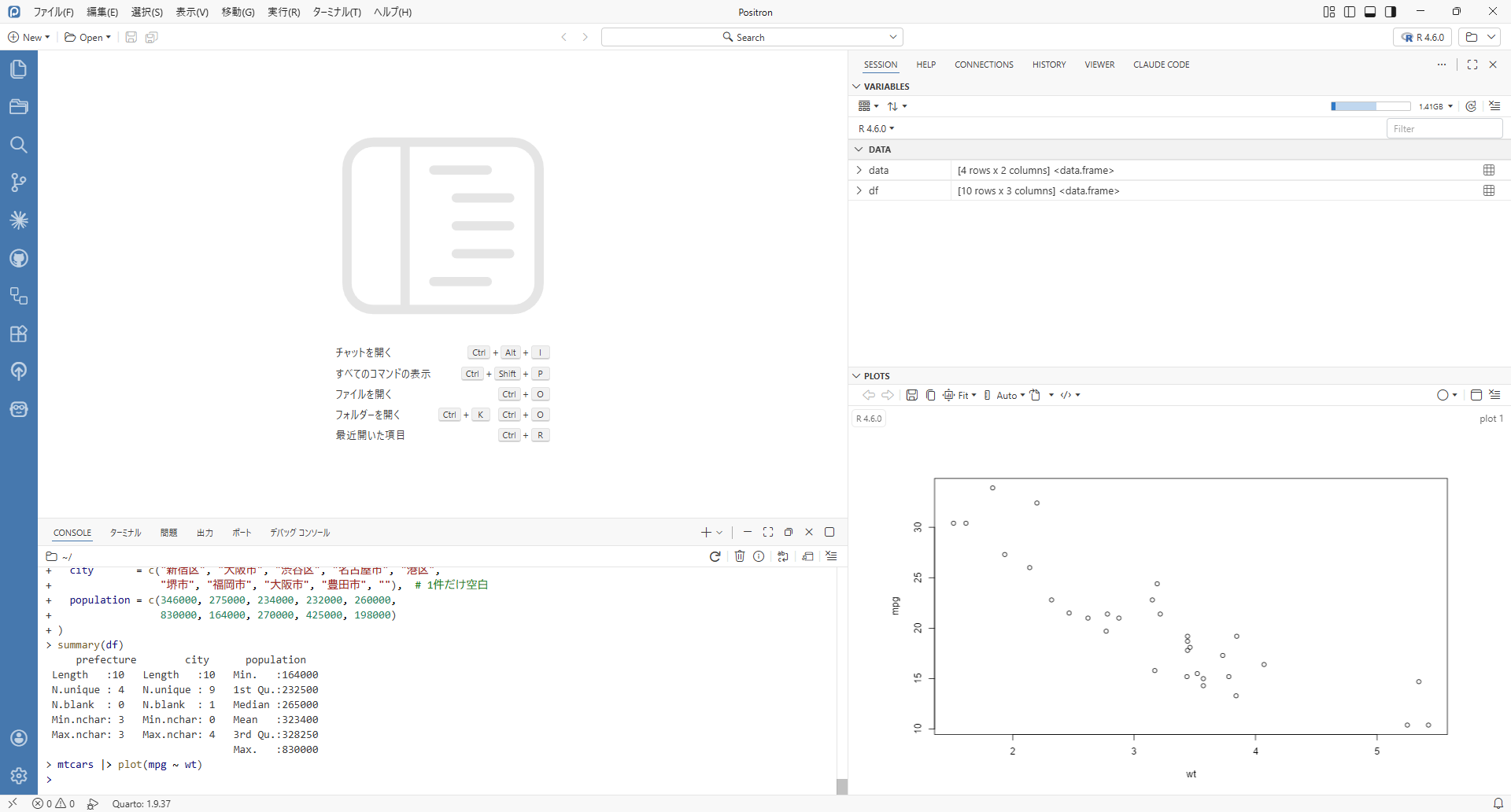
R 4.6.0 (“Because it was There”) is set for release on April 24th 2026. Here we summarise some of the more interesting changes that have been introduced. In previous blog posts, we have discussed the new features introduced in R 4.5.0 and earlier versions (see the links at the end of this post). Once R 4.6.0 is released, the full changelog will be available at the r-release ‘NEWS’ page. If you want to keep up to date with developments in base R, have a look at the r-devel ‘NEWS’ page. Data comes in all shapes and sizes. It can often be difficult to know where to start. Whatever your problem, Jumping Rivers can help. ! (values %in% collection) Code should be readable, and easily understood. And although it isn’t a natural language, there’s something off about code that reads like: If not a blog post is readable, I close the browser tab. To check if one (or more) value is in some collection, R has the %in% operator: "a" %in% letters [1] TRUE "a" in LETTERS [1] FALSE This is different from the in keyword, which you use when iterating over a collection: for (x in letters[1:3]) { message(x) } # a # b # c Sometimes you want to know whether a value is absent from a collection. The standard way to do this is to invert results from %in%: ! "a" %in% LETTERS [1] TRUE It’s unambiguous to the R interpreter. But it can be hard to read and understand - on scanning that statement, you might forget that ! acts after the %in%. As such, we often wrap the %in% expression with parentheses to make the code more clear: ! ("a" %in% LETTERS) [1] TRUE For the sake of clarity, many developers have implemented their own absence-checking operator. Writing a custom operator in R uses similar syntax to that used when writing a function: `%NOTIN%` = function(x, y) { ! (x %in% y) } "a" %NOTIN% LETTERS [1] TRUE Were you to write the same code multiple times in the same project, you would write a function. Similarly, if you (or your team) wrote the same function in multiple files or projects, you might add it to a package and import it. So if lots of package developers have implemented the same operator or function, across their CRAN packages, maybe it should be pushed to a higher plane… That is what has happened with the introduction of %notin% in R 4.6.0. An operator that was found across lots of separate packages has been moved up into base R: "a" %notin% LETTERS [1] TRUE "a" %notin% letters [1] FALSE DOI citations If you use R in your publications or your projects, you may need to provide a citation for it. rOpenSci has a blog post about citing R and R packages - why, when and how to do it. For the R project as a whole, there is a simple function citation() that provides the information you need: citation() To cite R in publications use: R Core Team (2026). _R: A Language and Environment for Statistical Computing_. R Foundation for Statistical Computing, Vienna, Austria. doi:10.32614/R.manuals . . A BibTeX entry for LaTeX users is @Manual{, ... } ... In R 4.6.0, a DOI (Digital Object Identifier) has been added to the citation entry to make it easier to reference R in your published work. summary(character_vector, character.method = "factor") str() and summary() are two of the first functions I reach for when exploring a dataset. For a data-frame, these tell me what type of columns are present (str(): chr, num, Date, …); and what is present in each column (summary(): gives the min, max, mean of each numeric column, for example). summary() works with data-structures other than just data-frames. For factors, summary() tells you how many observations of the factor levels were observed: # 'species' and 'island' are factor columns in `penguins` summary(penguins[1:3]) species island bill_len Adelie :152 Biscoe :168 Min. :32.10 Chinstrap: 68 Dream :124 1st Qu.:39.23 Gentoo :124 Torgersen: 52 Median :44.45 Mean :43.92 3rd Qu.:48.50 Max. :59.60 NA's :2 For character columns, the output from summary() has been a little obtuse: # R 4.5.0 # 'studyName' and 'Species' are character columns in `penguins_raw` summary(penguins_raw[1:3]) studyName Sample Number Species Length:344 Min. : 1.00 Length:344 Class :character 1st Qu.: 29.00 Class :character Mode :character Median : 58.00 Mode :character Mean : 63.15 3rd Qu.: 95.25 Max. :152.00 R 4.6.0 adds a neater way to summarise character vectors/columns to summary(): # R 4.6.0 summary(penguins_raw[1:3]) studyName Sample Number Species Length :344 Min. : 1.00 Length :344 N.unique : 3 1st Qu.: 29.00 N.unique : 3 N.blank : 0 Median : 58.00 N.blank : 0 Min.nchar: 7 Mean : 63.15 Min.nchar: 33 Max.nchar: 7 3rd Qu.: 95.25 Max.nchar: 41 Max. :152.00 We can also summarise character vectors/columns as if they were factors: # R 4.6.0 summary(penguins_raw[1:3], character.method = "factor") studyName Sample Number Species PAL0708:110 Min. : 1.00 Adelie Penguin (Pygoscelis adeliae) :152 PAL0809:114 1st Qu.: 29.00 Chinstrap penguin (Pygoscelis antarctica): 68 PAL0910:120 Median : 58.00 Gentoo penguin (Pygoscelis papua) :124 Mean : 63.15 3rd Qu.: 95.25 Max. :152.00 list.files(..., fixed = TRUE) Suppose we have three files in my working directory: abc.Rmd, CONTRIBUTING.md and README.md. If I want to obtain the filenames for the “.md” files from an R script, I can list those files that match a pattern: # R 4.5.0 list.files(pattern = ".md") [1] "abc.Rmd" "CONTRIBUTING.md" "README.md" Hmmm. In the pattern, . actually matches any character. In R 4.5.0, if I want to match the . character explicitly, I can escape it in the pattern. But pattern matching can lead to complicated code in R, because some characters are treated specially by the pattern matcher, and some are treated specially by R’s string parser. To tell R to find files with a literal ‘.md’ in the filename, we escape the . character twice: once for the . (to tell the pattern matcher to match a ., rather than any character), and once to escape the \ (to tell R that the subsequent \ is really a backslash) # R 4.5.0 list.files(pattern = "\.md") [1] "CONTRIBUTING.md" "README.md" R 4.0.x cleaned that up a bit, we can now use ‘raw strings’ in R. Everything between the parentheses in the next pattern is passed directly to the pattern matcher: # R 4.5.0 list.files(pattern = r"(\.md)") Now, in R 4.6.0, we can indicate to list.files() (and the synonym dir()) that our pattern is a fixed string (rather than a regular expression). With this, we don’t need to escape the . character to match the “.md” suffix. list.files(pattern = ".md", fixed = TRUE) [1] "CONTRIBUTING.md" "README.md" Other matters read.dcf() now allows comment lines, which means you can annotate your config files (e.g., for {lintr}). df |> plot(col2 ~ col1) can now be used for base plotting; this is a little neater for exploratory work than df |> plot(col2 ~ col1, data = _) C++20 is now the default C++ standard Trying out R 4.6.0 To take away the pain of installing the latest development version of R, you can use docker. To use the devel version of R, you can use the following commands: docker pull rstudio/r-base:devel-jammy docker run --rm -it rstudio/r-base:devel-jammy Once R 4.6 is the released version of R and the r-docker repository has been updated, you should use the following command to test out R 4.6. docker pull rstudio/r-base:4.6-jammy docker run --rm -it rstudio/r-base:4.6-jammy An alternative way to install multiple versions of R on the same machine is using rig. See also The R 4.x versions have introduced a wealth of interesting changes. These have been summarised in our earlier blog posts: R 4.0.0 R 4.1.0 R 4.2.0 R 4.3.0 R 4.4.0 R 4.5.0 For updates and revisions to this article, see the original post
r-bloggers.com